Conformational Variants of Duplex DNA Correlated with Cytosine-rich Chromosomal Fragile Sites
Albert G. Tsai, Aaron E. Engelhart, Ma’mon M. Hatmal, Sabrina I. Houston, Nicholas V. Hud, Ian S. Haworth, and Michael R. Lieber
Journal of Biological Chemistry, 2009, 284, pp 7157-7164. doi: 10.1074/jbc.M806866200
Publisher | ResearchGate | PubMed | Google Scholar
Scientific Abstract
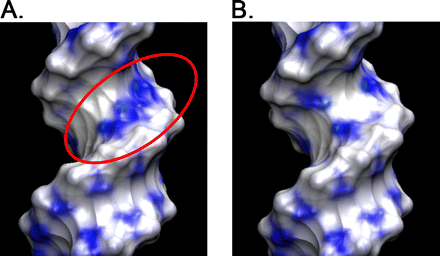
We found that several major chromosomal fragile sites in human lymphomas, including the bcl-2 major breakpoint region, bcl-1 major translocation cluster, and c-Myc exon 1-intron 1 boundary, contain distinctive sequences of consecutive cytosines exhibiting a high degree of reactivity with the structure-specific chemical probe bisulfite. To assess the inherent structural variability of duplex DNA in these regions and to determine the range of structures reactive to bisulfite, we have performed bisulfite probing on genomic DNA in vitro and in situ; on duplex DNA in supercoiled and linearized plasmids; and on oligonucleotide DNA/DNA and DNA/2′-O-methyl RNA duplexes. Bisulfite is significantly more reactive at the frayed ends of DNA duplexes, which is expected given that bisulfite is an established probe of single-stranded DNA. We observed that bisulfite also distinguishes between more subtle sequence/structural differences in duplex DNA. Supercoiled plasmids are more reactive than linear DNA; and sequences containing consecutive cytosines, namely GGGCCC, are more reactive than those with alternating guanine and cytosine, namely GCGCGC. Circular dichroism and x-ray crystallography show that the GGGCCC sequence forms an intermediate B/A structure. Molecular dynamics simulations also predict an intermediate B/A structure for this sequence, and probe calculations suggest greater bisulfite accessibility of cytosine bases in the intermediate B/A structure over canonical B- or A-form DNA. Electrostatic calculations reveal that consecutive cytosine bases create electropositive patches in the major groove, predicting enhanced localization of the bisulfite anion at homo-C tracts over alternating G/C sequences. These characteristics of homo-C tracts in duplex DNA may be associated with DNA-protein interactions in vivo that predispose certain genomic regions to chromosomal fragility.
Scientific Abstract
In this work, we found that certain sequences of DNA – notably, those associated with some cancers – exhibited unusual structures and a tendency to bind certain ions more tightly. These sequences form a slightly different double helix than regular DNA, which is more like the double helix formed by RNA. Computer simulations of these sequences suggested an explanation why these sequences bind the ions we examined more tightly. These characteristics may be relevant in explaining why the DNA sequences we examined are important in the development of these cancers.