Publications
Full list:
2026 Link to heading
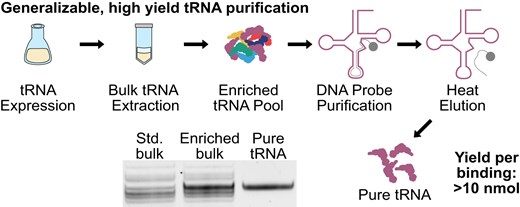
Purification of post-transcriptionally modified tRNAs for enhanced cell-free translation systems
Kalb, Evan M; Alejo, Jose L; Dias-Fields, Leticia; Knudson, Isaac; Davisson, Joshua A; Maldonado, Efren; Chattrakun, Kanokporn; Lin, Shangsi; Lee, Jung Yeon; He, Tianchen;
Nucleic Acids Research (2026) · doi:10.1093/nar/gkag208
2025 Link to heading
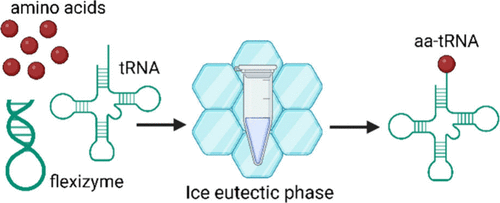
High yield, low magnesium flexizyme reactions in a water-ice eutectic phase
Joshua Davisson, Jose Alejo, Mace Blank, Evan Kalb, Angelin Prasad, Isaac Knudson, Alanna Schepartz, Aaron E Engelhart, Katarzyna P Adamala
Biochemistry (2025) · doi: 10.1021/acs.biochem.5c00371
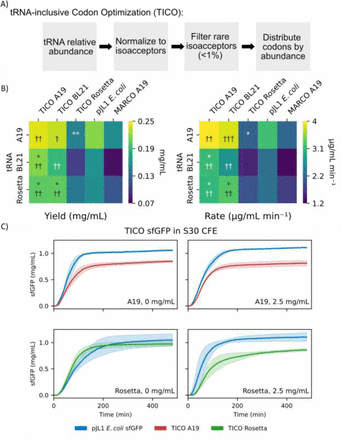
An Expanded Repertoire of tRNA Sources for Cell-Free Protein Synthesis
Kalb, Evan M; Vincent, Russel M; Engelhart, Aaron E; Church, George M; Adamala, Katarzyna P;
bioRxiv (2025) · doi:10.1101/2025.08.20.671396
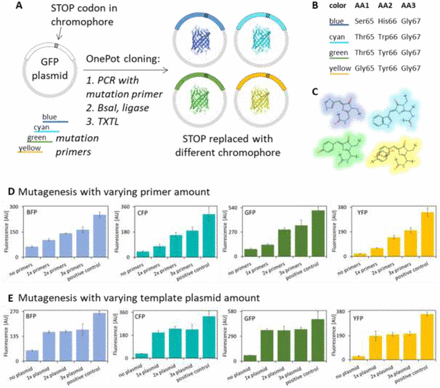
One-pot cloning and protein expression platform for genetic engineering
Sato, Wakana; Sharon, Judee; Cash, Brock; Deich, Christopher; Gaut, Nathaniel J; Heili, Joseph; Engelhart, Aaron E; Adamala, Katarzyna P;
bioRxiv (2025) · doi:10.1101/2025.08.28.672974
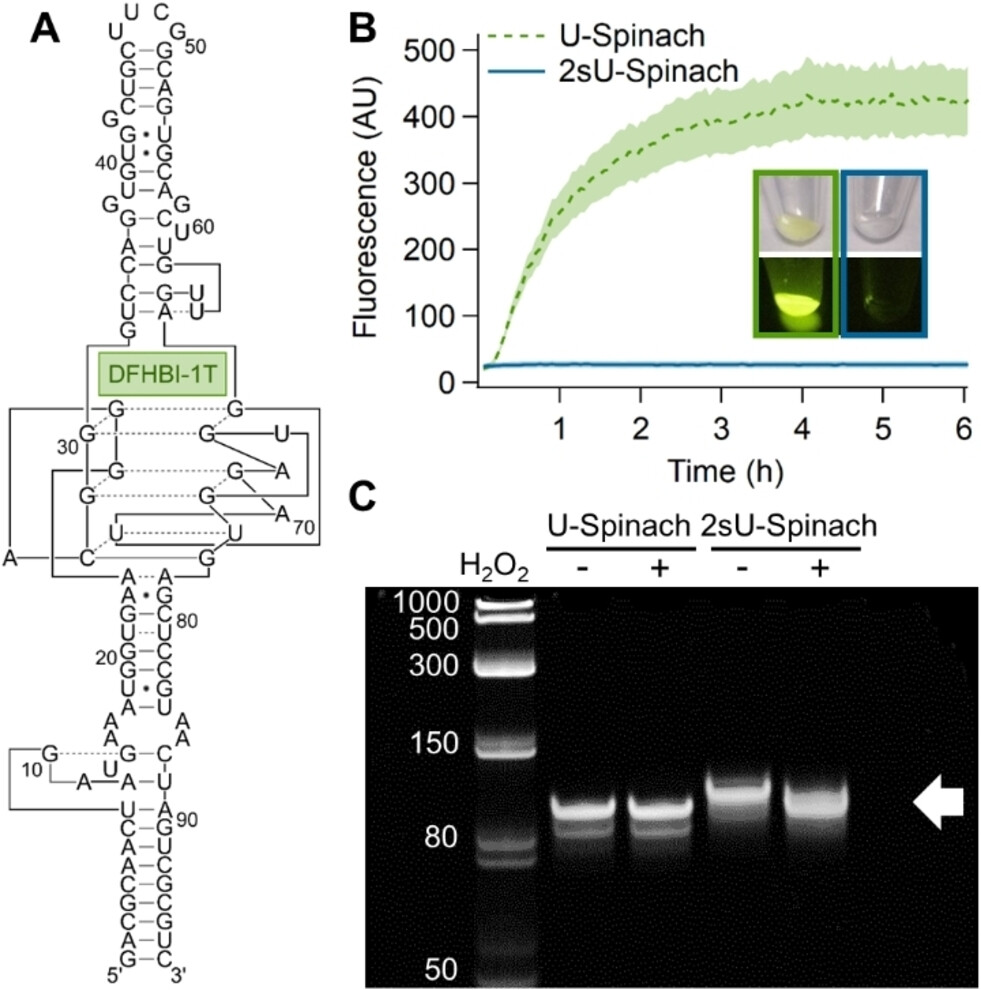
Activation of Caged Functional RNAs by An Oxidative Transformation
Heili JM, Adamala KP, Engelhart AE.
Chembiochem (2025) · doi:10.1002/cbic.202401056
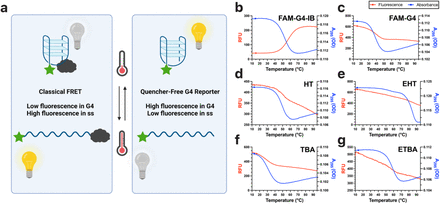
Quencher-Free Fluorescence Monitoring of G-Quadruplex Folding
Parada Z, Hoog TG, Adamala KP, Engelhart AE.
ACS Omega (2025) · doi:10.1101/2024.01.31.578026
2024 Link to heading
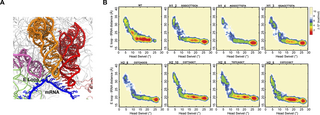

Alternate conformational trajectories in ribosome translocation
Alejo JL, Girodat D, Hammerling MJ, Willi JA, Jewett MC, Engelhart AE, Adamala KP.
PLoS Comput Biol (2024) · doi:10.1371/journal.pcbi.1012319

Emergent ribozyme behaviors in oxychlorine brines indicate a unique niche for molecular evolution on Mars
Tanner G. Hoog, Matthew R. Pawlak, Nathaniel J. Gaut, Gloria C. Baxter, Thomas A. Bethel, Katarzyna P. Adamala & Aaron E. Engelhart
Nature Communications (2024) · doi:10.1038/s41467-024-48037-2
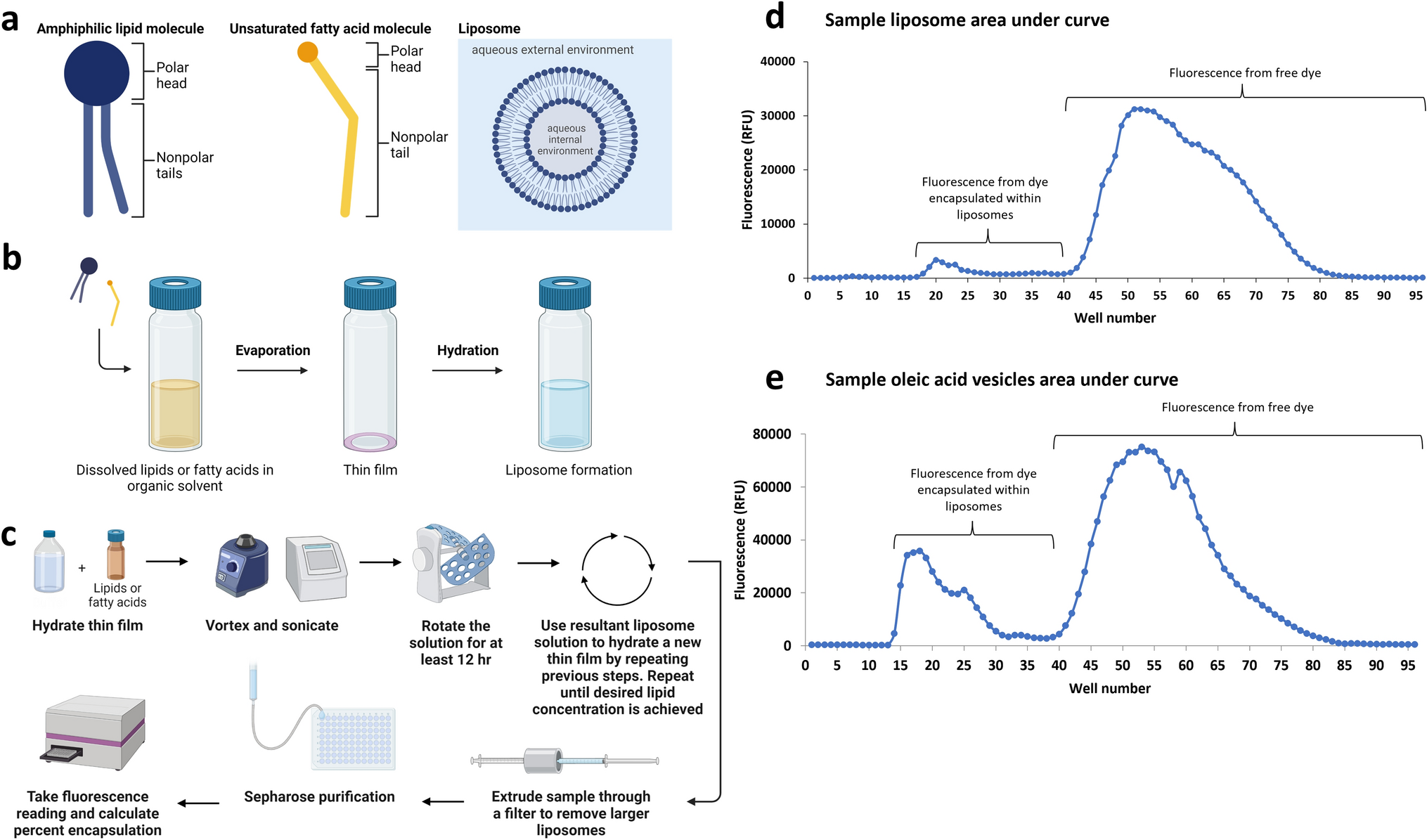
Sequential gentle hydration increases encapsulation in model protocells
EM Gehlbach, AO Robinson, AE Engelhart, KP Adamala
Discover Life (2024) · doi:10.1007/s11084-024-09645-6
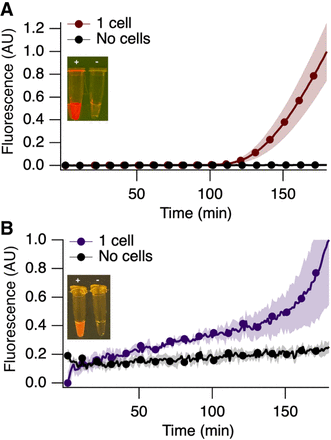
PACRAT: Pathogen detection with aptamer-observed cascaded recombinase polymerase amplification-in vitro transcription
Pavana Khan, Lauren Aufdembrink, Katarzyna P Adamala, Aaron Engelhart
RNA (2024) · doi:10.1261/rna.079891.123
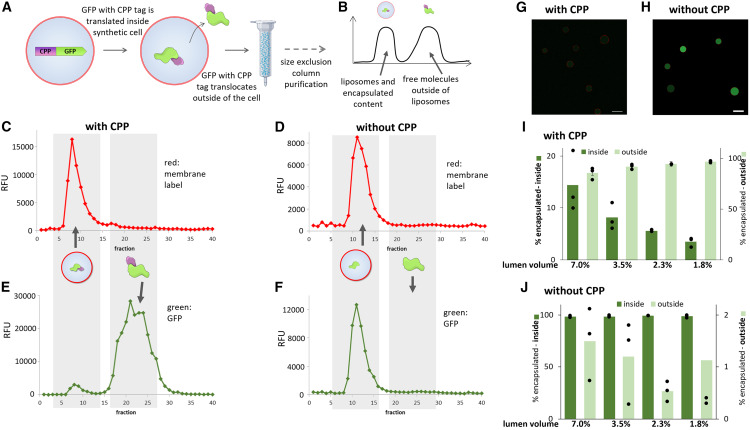
Controlled exchange of protein and nucleic acid signals from and between synthetic minimal cells
Joseph M Heili, Kaitlin Stokes, Nathaniel J Gaut, Christopher Deich, Jose Gomez-Garcia, Brock Cash, Matthew R Pawlak, Aaron E Engelhart, Katarzyna P Adamala
Cell Systems (2024) · doi:10.1016/j.cels.2023.12.008
2023 Link to heading
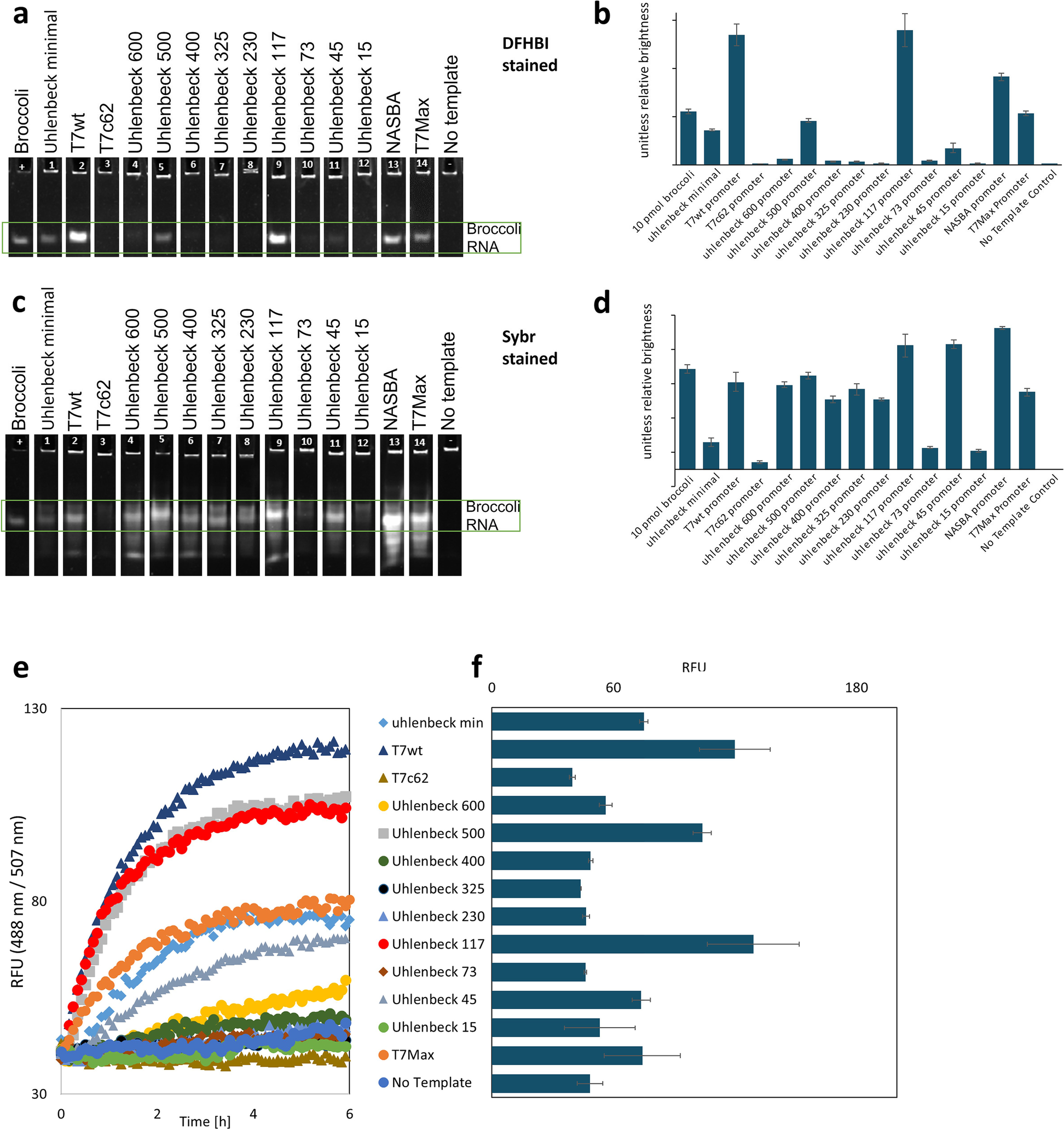
T7Max transcription system
Christopher Deich, Brock Cash, Wakana Sato, Judee Sharon, Lauren Aufdembrink, Nathaniel J Gaut, Joseph Heili, Kaitlin Stokes, Aaron E Engelhart, Katarzyna P Adamala
J Biol Eng (2023) · doi:10.1186/s13036-023-00323-1
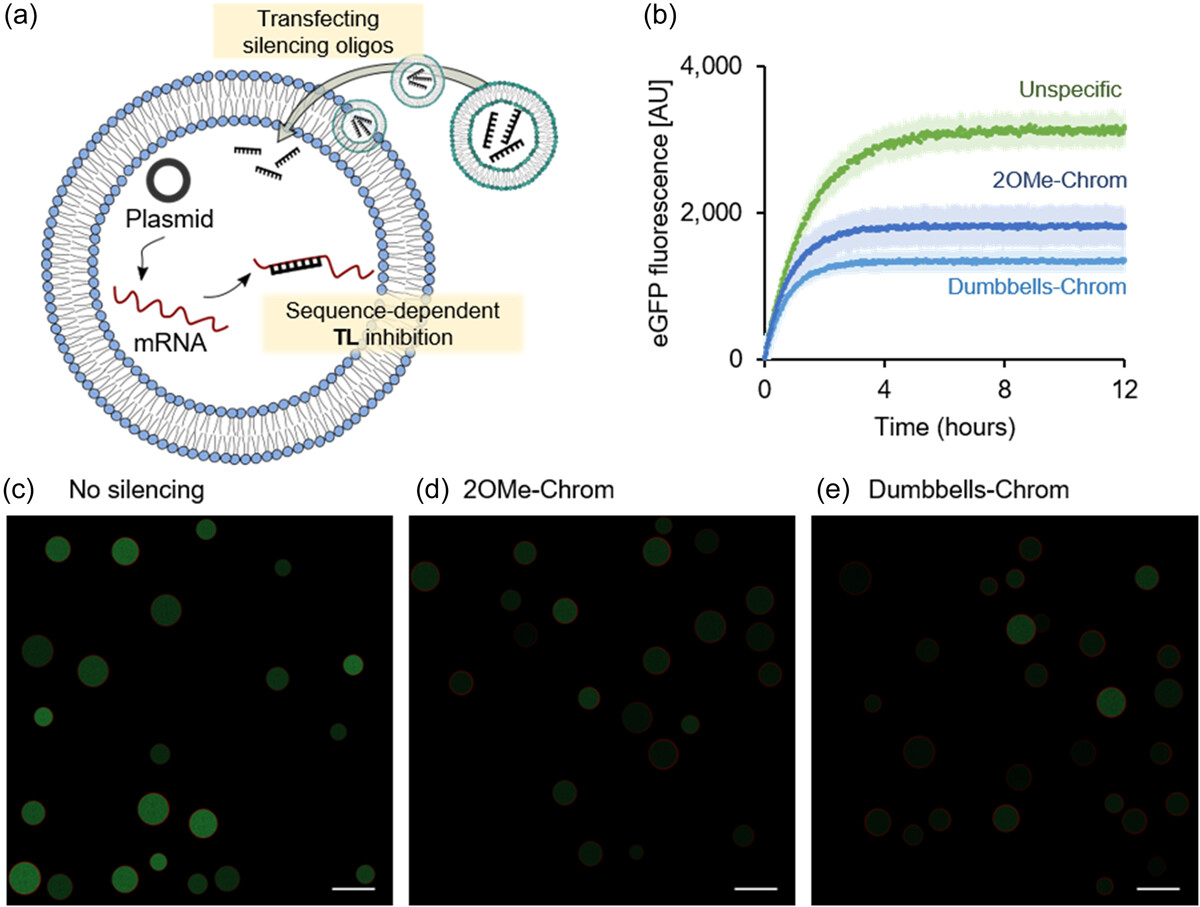
A gene expression control technology for cell‐free systems and synthetic cells via targeted gene silencing and transfection
Wakana Sato, Melanie Rasmussen, Nathaniel Gaut, Mahima Devarajan, Kaitlin Stokes, Christopher Deich, Aaron E Engelhart, Katarzyna P Adamala
Biotechnology and bioengineering (2023) · doi:10.1002/bit.28422
New Aequorea Fluorescent Proteins for Cell-Free Bioengineering
C Deich, NJ Gaut, W Sato, AE Engelhart, KP Adamala
ACS Synthetic Biology (2023) · doi:10.1021/acssynbio.3c00057
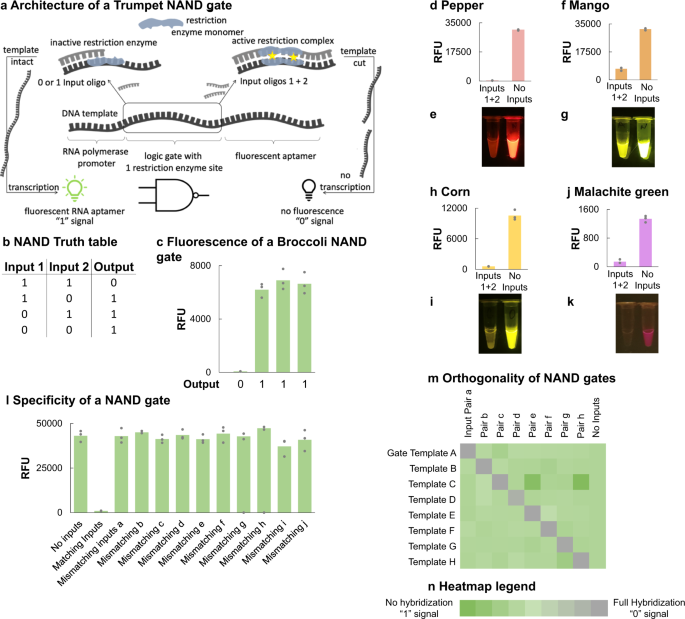
Trumpet is an operating system for simple and robust cell-free biocomputing
Judee A. Sharon, Chelsea Dasrath, Aiden Fujiwara, Alessandro Snyder, Mace Blank, Sam O’Brien, Lauren M. Aufdembrink, Aaron E. Engelhart & Katarzyna P. Adamala
Nature communications (2023) · doi:10.1038/s41467-023-37752-x
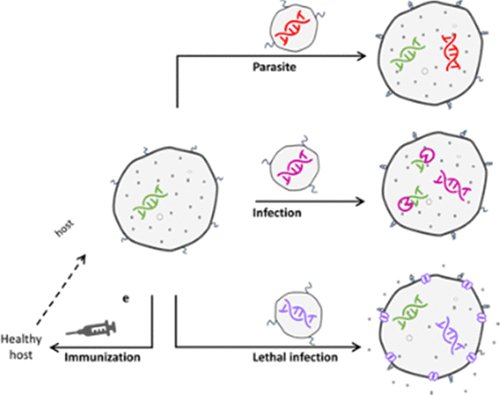
Parasites, infections and inoculation in synthetic minimal cells
Brock Cash, Nathaniel J. Gaut, Christopher Deich, Laura L. Johnson, Aaron E. Engelhart, Katarzyna P. Adamala
ACS Omega (2023) · doi:10.1021/acsomega.2c07911
2022 Link to heading
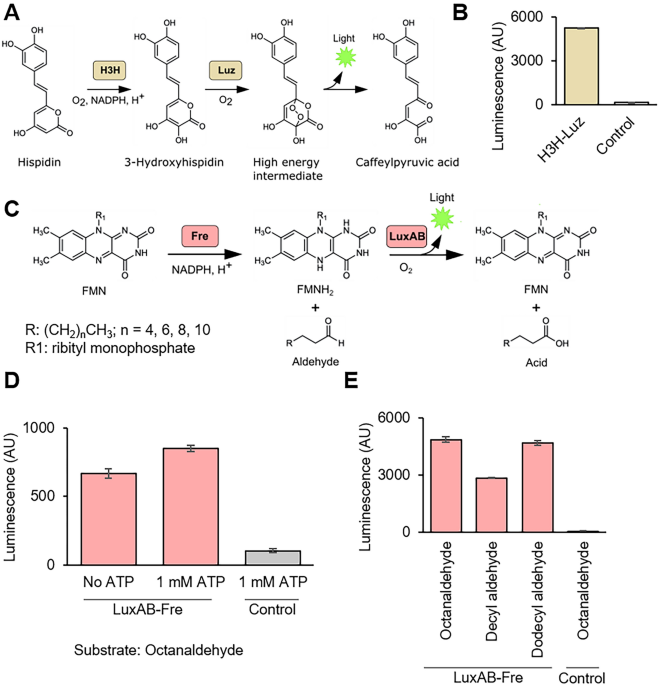
Expanding luciferase reporter systems for cell-free protein expression
Wakana Sato, Melanie Rasmussen, Christopher Deich, Aaron E Engelhart, Katarzyna P Adamala
Sci Rep (2022) · doi:10.1038/s41598-022-15624-6
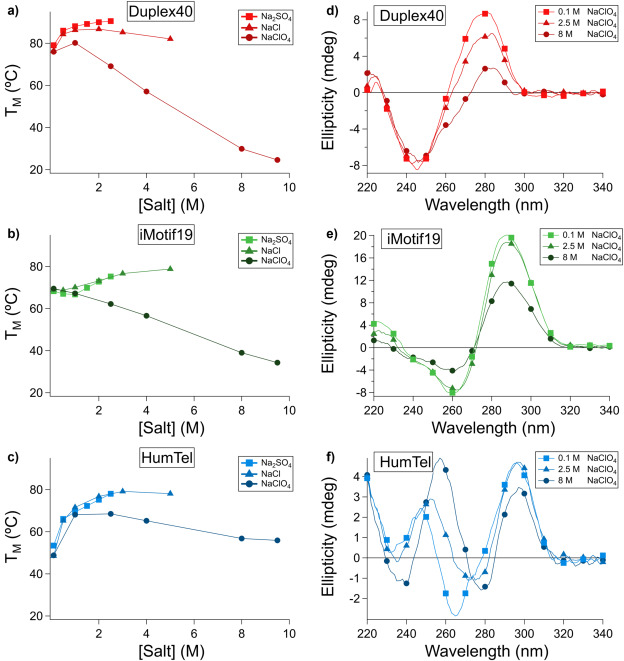
DNA G-quadruplexes are uniquely stable in the presence of denaturants and monovalent cations
Tanner G Hoog, Matthew R Pawlak, Benjamin F Bachan, Aaron E Engelhart
Biochemistry and Biophysics Reports (2022) · doi:10.1016/j.bbrep.2022.101238
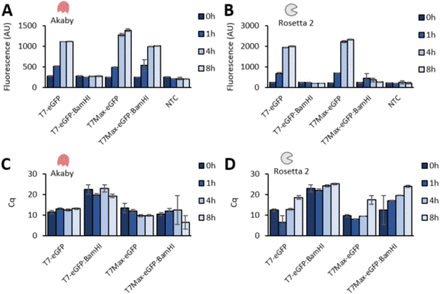
Akaby-Cell-free protein expression system for linear templates
Sato W, Sharon J, Deich C, Gaut N, Cash B, Engelhart AE, Adamala KP.
PLoS One (2022) · doi:10.1101/2021.11.03.467179
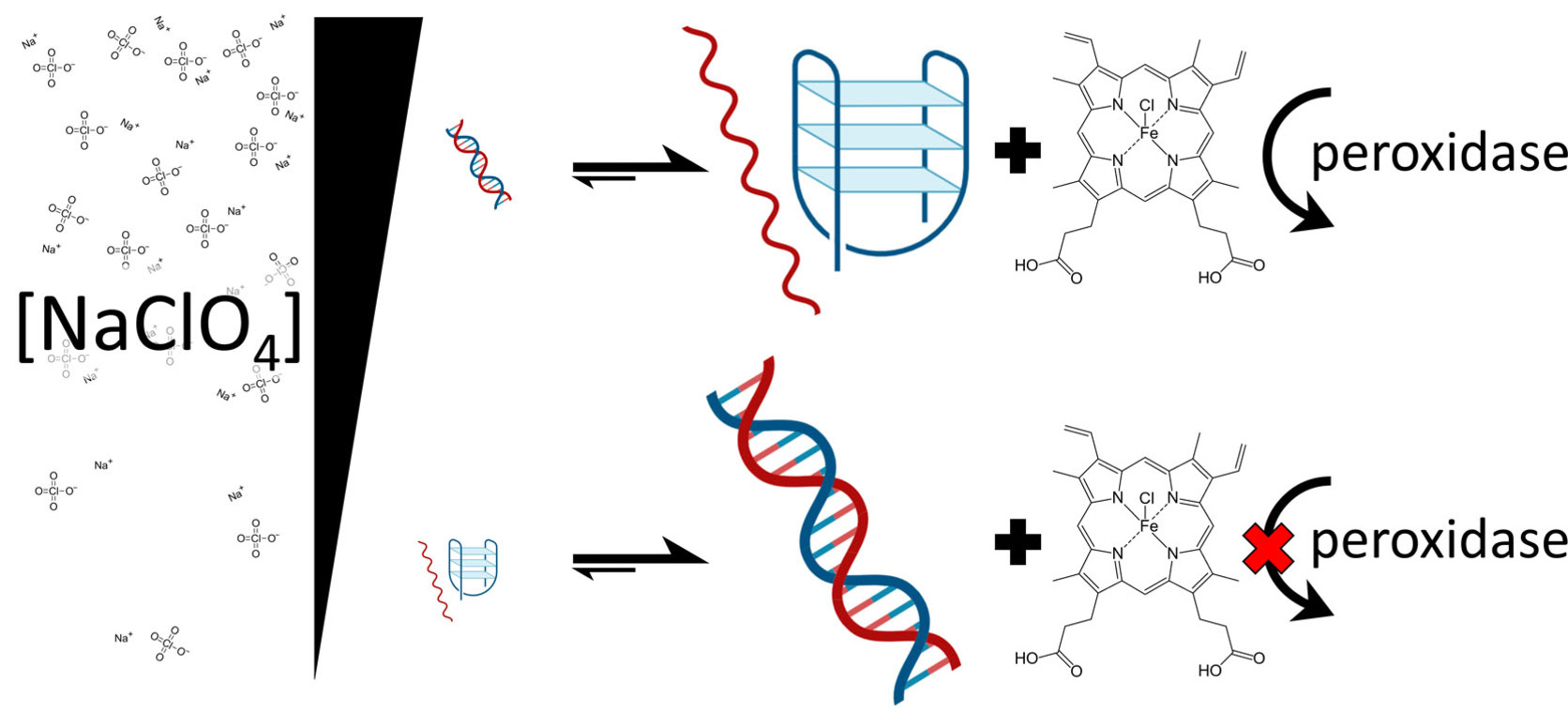
Switchable DNA-Based Peroxidases Controlled by a Chaotropic Ion
Tanner G Hoog, Matthew R Pawlak, Lauren M Aufdembrink, Benjamin R Bachan, Matthew B Galles, Nicholas B Bense, Katarzyna P Adamala, Aaron E Engelhart
ChemBioChem (2022) · doi:10.1002/cbic.202200090
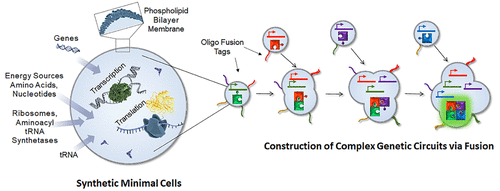
Programmable Fusion and Differentiation of Synthetic Minimal Cells
Nathaniel J Gaut, Jose Gomez-Garcia, Joseph M Heili, Brock Cash, Qiyuan Han, Aaron E Engelhart, Katarzyna P Adamala
ACS Synth. Biol (2022) · doi:10.1021/acssynbio.1c00519
2021 Link to heading

Astrobiology on habitable worlds: The case for considering prebiotic chemistry in mission design
Aaron Engelhart, Jennifer G Blank, Christopher Carr, Henderson James Cleaves, and Kennda Lynch
Planetary Science and Astrobiology Decadal Survey 2023-2032 (2021) · doi:10.3847/25c2cfeb.52e0f294
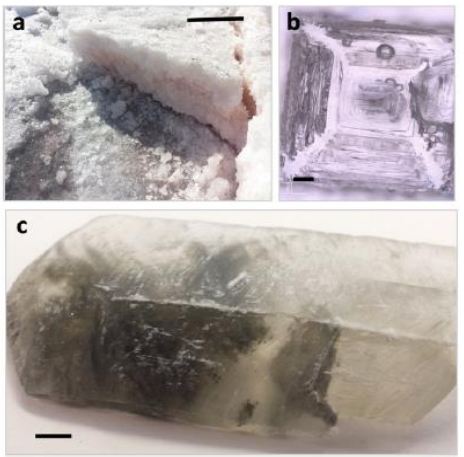

Salty Environments: The importance of evaporites and brine environments as habitats and preservers of biosignatures
Scott Perl, Solmaz Adeli, Chhandak Basu, Bonnie K. Baxter, Jeff Bowman, Eric Boyd, Morgan Cable, Aaron J. Celestian, Charles S. Cockell, Frank A. Corsetti, Kate L. Craft, Aaron Engelhart, Alberto G. Fairen, Suniti Karunatillake, Kennda Lynch, Sally Potter-McIntyre, Frances Rivera-Hernandez, Mark Schneegurt, Susanne Schwenzer, Svetlana Shkolyar, Bethany Theiling, Brian Wade, and Jon Zaloumis
Planetary Science and Astrobiology Decadal Survey 2023-2032 (2021) · doi:10.3847/25c2cfeb.d0fffbba
2020 Link to heading
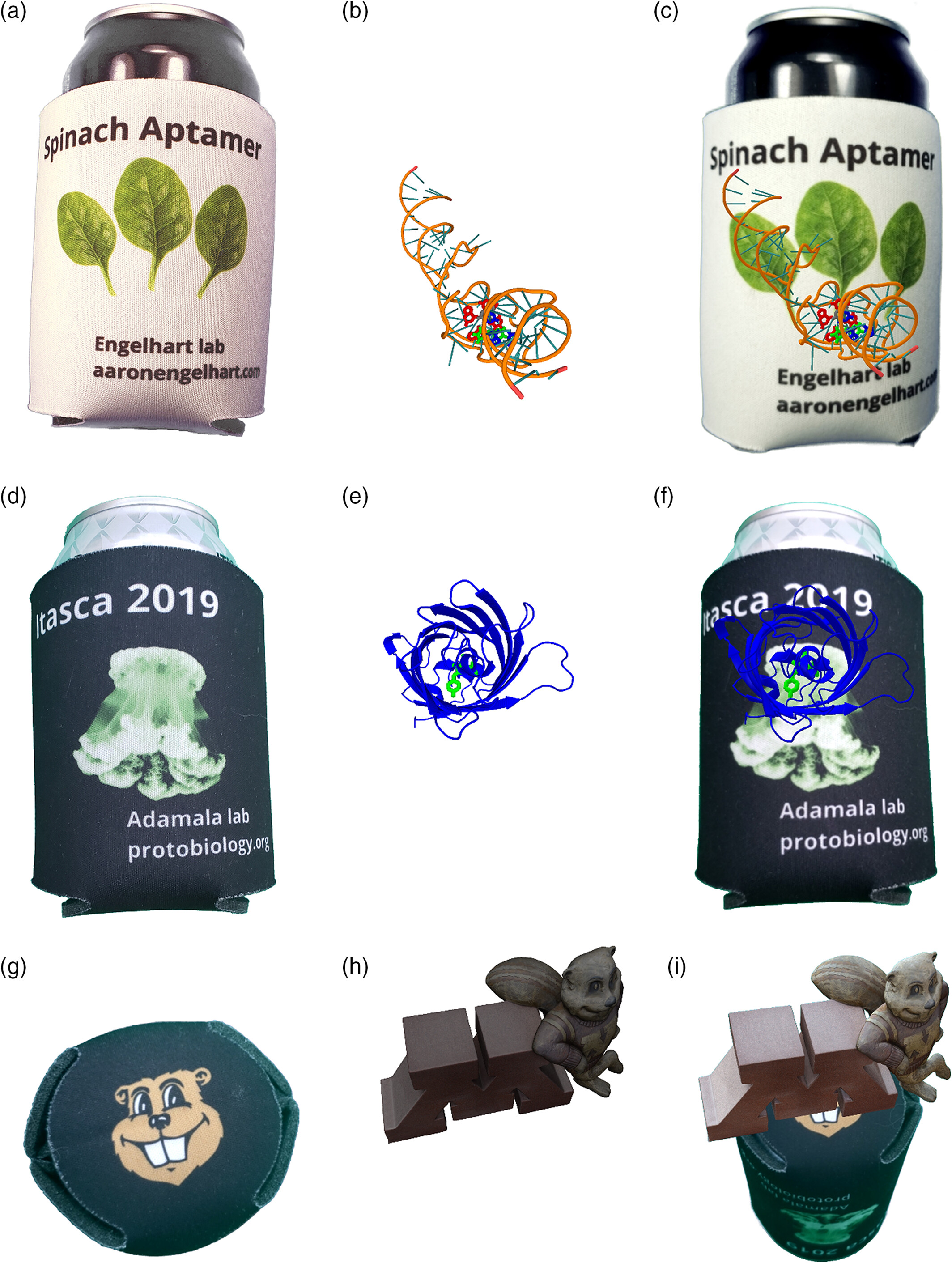

Rapid deployment of smartphone‐based augmented reality tools for field and online education in structural biology
Tanner G Hoog, Lauren M Aufdembrink, Nathaniel J Gaut, Rou‐Jia Sung, Katarzyna P Adamala, Aaron E Engelhart
Biochemistry and Molecular Biology Education (2020) · doi:10.1002/bmb.21396
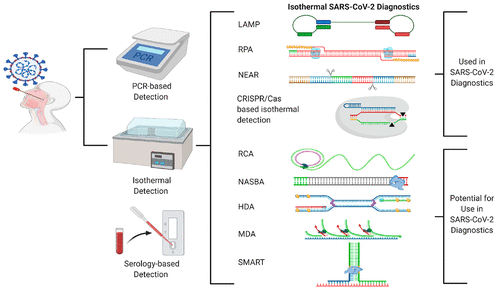
Isothermal SARS-CoV-2 diagnostics: tools for enabling distributed pandemic testing as a means of supporting safe reopenings
Pavana Khan, Lauren M Aufdembrink, Aaron E Engelhart
ACS Synth. Biol (2020) · doi:10.1021/acssynbio.0c00359
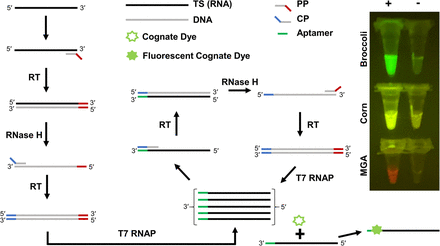
Highly specific, multiplexed isothermal pathogen detection with fluorescent aptamer readout
Lauren M Aufdembrink, Pavana Khan, Nathaniel J Gaut, Katarzyna P Adamala, Aaron E Engelhart
RNA (2020) · doi:10.1261/rna.075192.120
2019 Link to heading
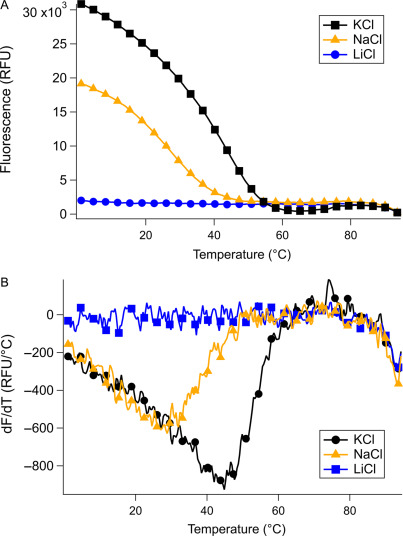
Methods for thermal denaturation studies of nucleic acids in complex with fluorogenic dyes
Lauren M Aufdembrink, Tanner G Hoog, Matthew R Pawlak, Benjamin F Bachan, Joseph M Heili, Aaron E Engelhart
Methods in Enzymology (2019) · doi:10.1016/bs.mie.2019.05.029
2018 Link to heading
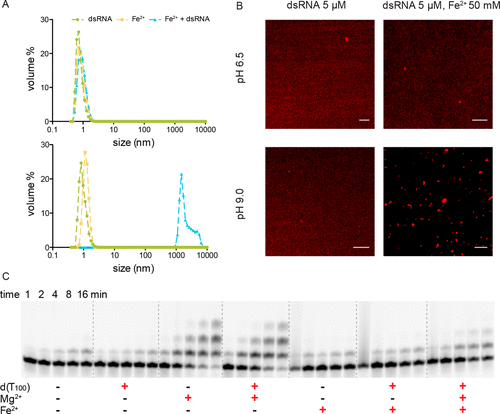
Catalysis of template-directed nonenzymatic RNA copying by Iron (II)
Lin Jin, Aaron E Engelhart, Weicheng Zhang, Katarzyna Adamala, Jack W Szostak
J. Am. Chem. Soc (2018) · doi:10.1021/jacs.8b09617
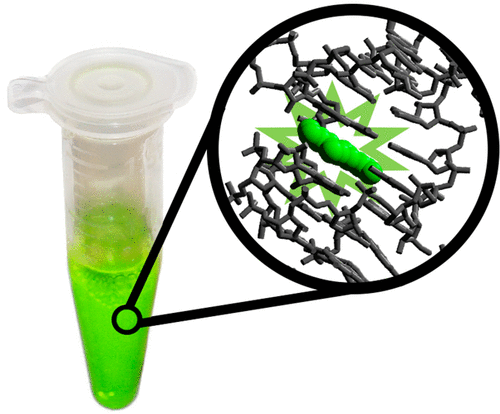
Real-Time Visualization of in Vitro Transcription of a Fluorescent RNA Aptamer: An Experiment for the Upper-Division Undergraduate or First-Year Graduate Laboratory
Joseph M Heili, Jose Gomez-Garcia, Nathaniel J Gaut, Brock W Cash, Lauren M Aufdembrink, Brent A Heffron, Joshua D Shirley, Erin E Carlson, Katarzyna P Adamala, Aaron E Engelhart
Journal of Chemical Education (2018) · doi:10.1021/acs.jchemed.7b00735
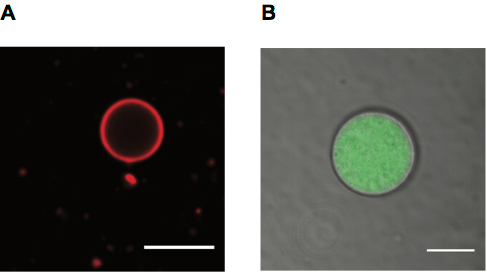
Preparation, Purification, and Use of Fatty Acid-containing Liposomes
Aaron E. Engelhart
Journal of Visualized Experiments (2018) · doi:10.3791/57324
2017 Link to heading
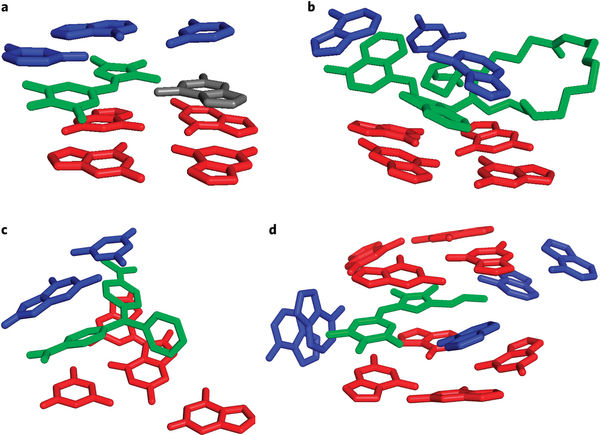
RNA imaging: A tale of two G-quadruplexes
Aaron E. Engelhart
Nature Chemical Biology (2017) · doi:10.1038/nchembio.2492
2016 Link to heading
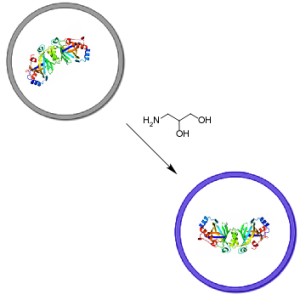
Collaboration between primitive cell membranes and soluble catalysts
Katarzyna P. Adamala, Aaron E. Engelhart (joint first author), and Jack W. Szostak
Nature Communications (2016) · doi:10.1038/ncomms11041
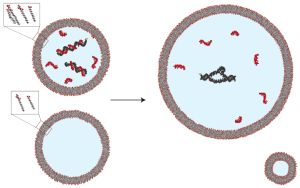
A simple physical mechanism enables homeostasis in primitive cells
Aaron E. Engelhart, Katarzyna P. Adamala, and Jack W. Szostak
Nature Chemistry (2016) · doi:10.1038/nchem.2475
2015 Link to heading
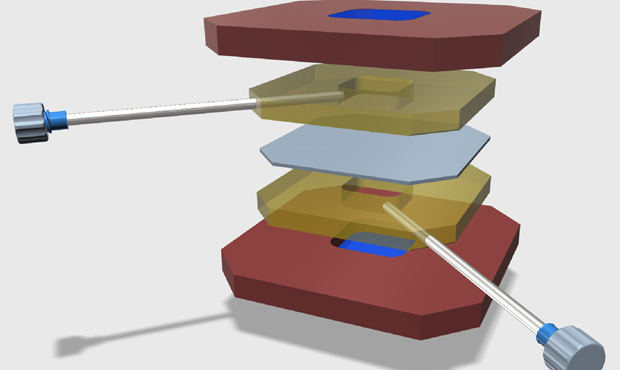
Construction of a liposome dialyzer for the preparation of high-value, small-volume liposome formulations
Nature Protocols (2015) · doi:10.1038/nprot.2015.054
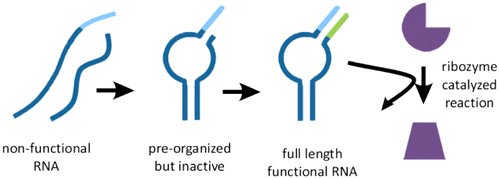
Generation of functional RNAs from inactive oligonucleotide complexes by non-enzymatic primer extension
JACS (2015) · doi:10.1021/ja511564d
2014 Link to heading
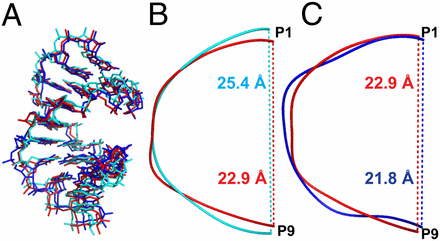
Structural insights into the effects of 2′-5′ linkages on the RNA duplex
PDB Entries: 4MS9 , 4MSB , 4MSR
PNAS (2014) · doi:10.1073/pnas.1317799111
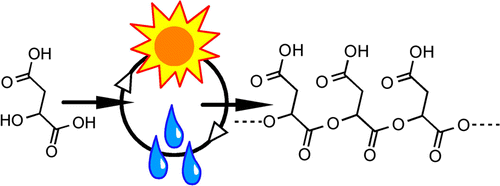
Ester formation and hydrolysis during wet–dry cycles: Generation of far-from-equilibrium polymers in a model prebiotic reaction
Macromolecules (2014) · doi:10.1021/ma402256d
2013 Link to heading
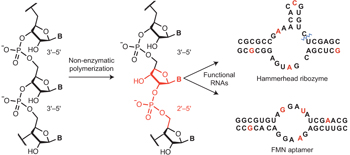
Functional RNAs exhibit tolerance for non-heritable 2′–5′ versus 3′–5′ backbone heterogeneity
Nature Chemistry (2013) · doi:10.1038/nchem.1623
2012 Link to heading
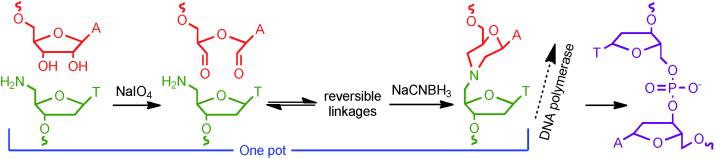
Nonenzymatic ligation of DNA with a reversible step and a final linkage that can be used in PCR
ChemBioChem (2012) · doi:10.1002/cbic.201200167
2011 Link to heading
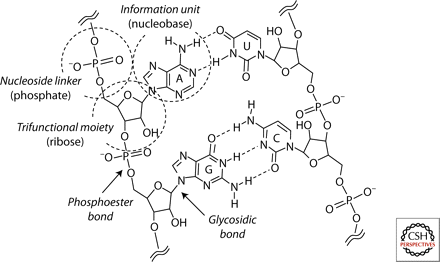
Primitive genetic polymers
Cold Spring Harbor Perspectives in Biology (2011) · doi:10.1101/cshperspect.a002196
2010 Link to heading

DNA and RNA in anhydrous media: duplex, triplex, and G-Quadruplex secondary structures in a deep eutectic solvent
Angewandte Chemie (2010) · doi:10.1002/ange.201001561
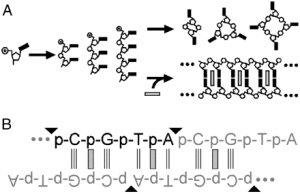
Intercalation as a means to suppress cyclization and promote polymerization of base-pairing oligonucleotides in a prebiotic world
PNAS (2010) · doi:10.1073/pnas.0914172107
2009 Link to heading
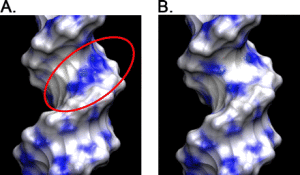
Conformational variants of duplex DNA correlated with cytosine-rich chromosomal fragile sites
J. Biol. Chem (2009) · doi:10.1074/jbc.m806866200

Evidence of strong hydrogen bonding by 8-aminoguanine
ChemComm (2009) · doi:10.1039/b818409g

Metal ion interactions with G-Quadruplex structures
RSC Book Chapter (2009) · doi:10.1039/9781847558763-00118

Sequence-Specific DNA-Metal Ion Interactions
RSC Book Chapter (2009) · doi:10.1039/9781847558763-00075
2008 Link to heading
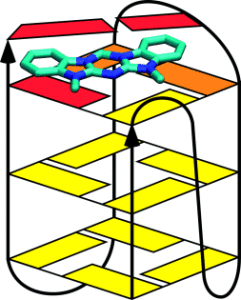
Submicromolar, selective G-Quadruplex ligands from one pot: thermodynamic and structural studies of human telomeric DNA binding by azacyanines
ChemBioChem (2008) · doi:10.1002/cbic.200800234